- HOME
- Deep purple smoke on the water released
- Nbc app download
- VENUE
- Windows 10 xbox 360 controller driver
- RSVP
- Free symantec endpoint protection 14 download full version
- REGISTRY
- CONTACT
- Best eml to pst converter
- Wwe games for wii
- Finding snp using bioedit
- Dil ko karaar aaya
- Battletech tactical operations torrent
- Copytrans manager torrent
- Lirik lagu a7x unholy
- Best total annihilation maps
- Wikipedia fairy tail episodes
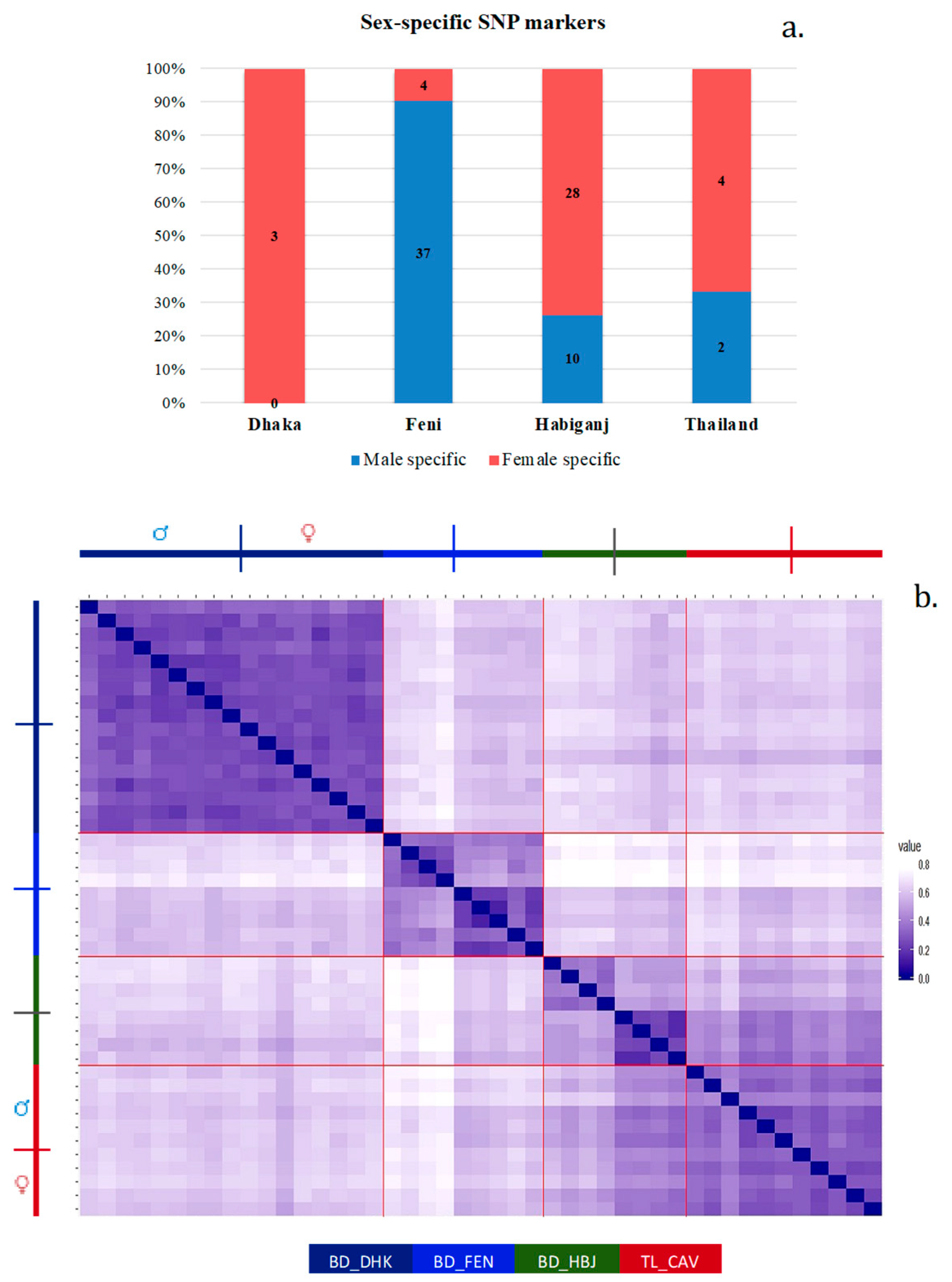
This is showing a comparison of the VCF file from the original SNP discovery analysis for one dataset, and a second run of the SNP discovery pipeline on the same dataset, but with technical replicates removed (blue). Personally, I can validate this by saying that I recently ran vcf-compare and bcftools isec (as per above) and used the results to generate a Venn diagram with jvenn, shown below: Once you had the output of running these programs in hand, it would then be possible to do a number of things, such as report common/different SNPs between runs or treatments, conduct statistical anlaysis, or create a Venn diagram of common/different SNPs between multiple VCF files to visualize the differences.
FINDING SNP USING BIOEDIT MAC
On mac or Linux with bcftools installed, you could use something like the following (where $ is the command line prompt) to get the list of SNPs at the intersection of two or more VCF files: $ bcftools isec -n +2 | bgzip -c > isec_file1-v-2_Īlternatively, if you wanted just statistics on the numbers of SNPs/variants or genotypes in common between files, you could use the vcf-compare tool that comes with vcftools.
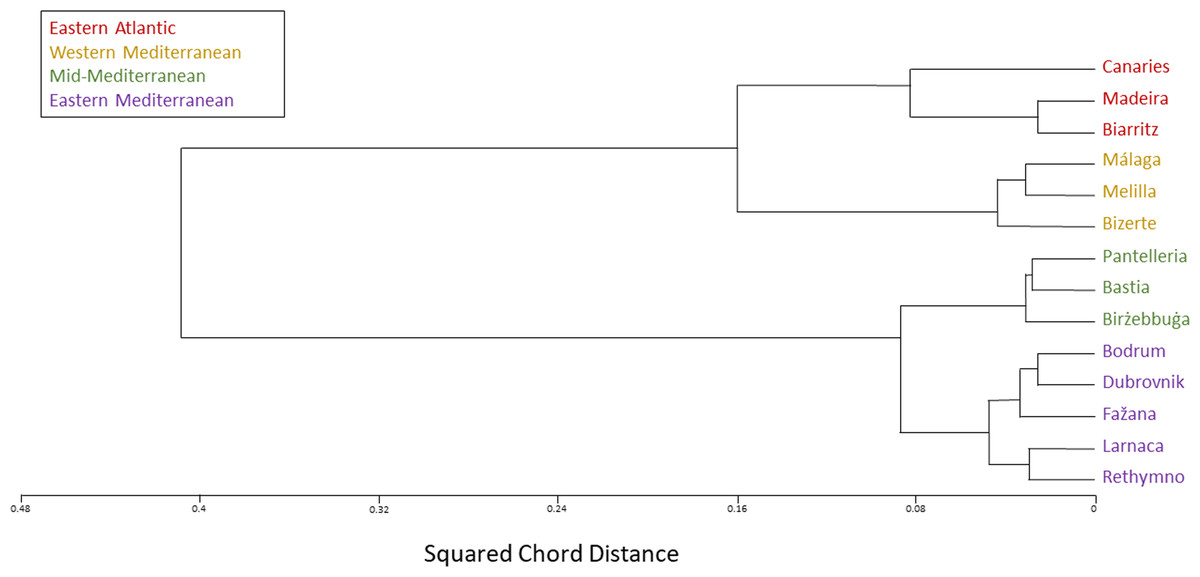
To mention other options, bcftools is supposedly faster at this, and if you use bcftools what you want is the intersection function, isec. The output file has the suffix “.diff.sites_in_files”.” “Outputs the sites that are common / unique to each file. This option for the -diff flag is listed in the documentation as having the following function: To mention other options, bcftools is supposedly faster at this, and if you use bcftools what you want is the intersection function, isec. Title: Methodology for SNP characterization at amino acid level.

cerevisiae S288c, version 12.0) was created using the SNP data produced for CEN.PK113-7D compared to S288c. from different SNP discovery pipelines, or two treatments of an experiment)?”, you might ask.īelow, I provide a post based on my recent answer to this ResearchGate question that provides some solutions for this problem.įirst, the vcftools -diff -diff-site option would work for this specific case. Description: The metabolic map, produced using the Saccharomyces Genome Database (SGD) Expression Viewer (SRI International Pathway Tools version 12.0, based upon S. “What are the SNPs or variants that are shared in common between two VCF files I created (e.g.